crested.pl.modisco.modisco_results#
- crested.pl.modisco.modisco_results(classes, contribution, contribution_dir, num_seq, viz='contrib', min_seqlets=0, verbose=False, y_min=-0.05, y_max=0.25, background=None, trim_pattern=True, trim_ic_threshold=0.1, **kwargs)#
Plot genomic contributions for the given classes.
Requires the modisco results to be present in the specified directory. The contribution scores are trimmed based on information content (IC).
- Parameters:
classes (
list[str]) – List of classes to plot genomic contributions for.contribution (
str) – Contribution type to plot. Choose either “positive” or “negative”.contribution_dir (
str) – Directory containing the modisco results. Each class should have a separate modisco .h5 results file in the format {class}_modisco_results.h5.num_seq (
int) – Total number of sequences used for the modisco run. Necessary to calculate the percentage of sequences with the pattern.viz (
str(default:'contrib')) – Visualization method. Choose either “contrib” or “pwm”.min_seqlets (
int(default:0)) – Minimum number of seqlets required for a pattern to be considered.verbose (
bool(default:False)) – Print verbose output.y_min (
float(default:-0.05)) – Minimum y-axis limit for the plot if viz is “contrib”.y_max (
float(default:0.25)) – Maximum y-axis limit for the plot if viz is “contrib”.background (
list[float] (default:None)) – Background probabilities for each nucleotide. Default is [0.27, 0.23, 0.23, 0.27].trim_pattern (
bool(default:True)) – Boolean for trimming modisco patterns.trim_ic_threshold (
float(default:0.1)) – If trimming patterns, indicate threshold.width – Width of the newly created figure. Default is 6*`len(classes)`.
height – Height of the newly created figure. Default is 2*`max_num_patterns`.
sharex – Whether to share the x axes of the created plots. Default is False.
sharey – Whether to share the y axes of the created plots. Default is False.
kwargs – Additional arguments passed to
render_plot()to control the final plot output. Please seerender_plot()for details.
- Return type:
Examples
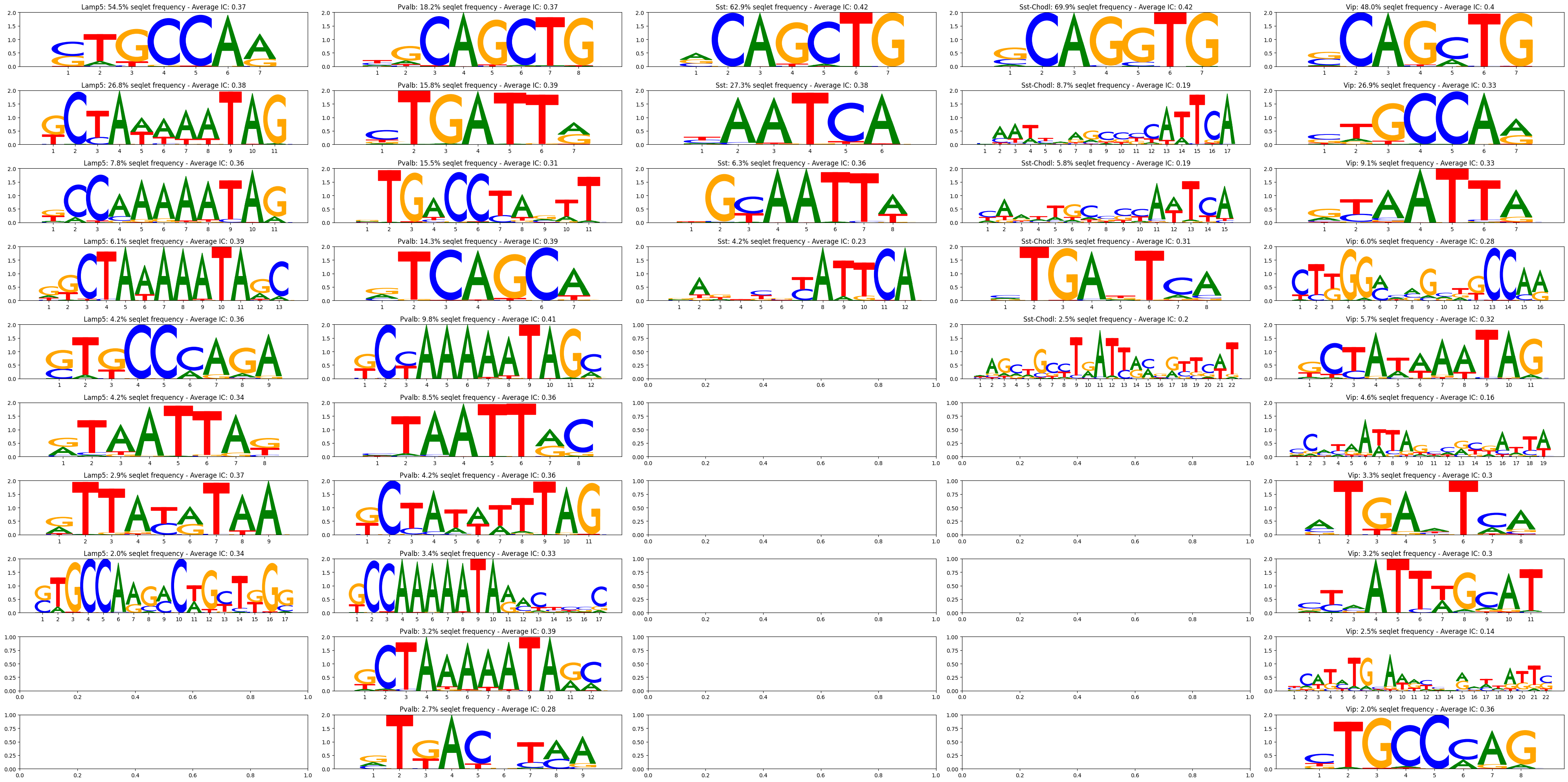
>>> crested.pl.modisco.modisco_results( ... classes=["Lamp5", "Pvalb", "Sst", ""Sst-Chodl", "Vip"], ... contribution="positive", ... contribution_dir="/path/to/modisco_results", ... num_seq=1000, ... viz="pwm", ... )